Introduction
Mexico is considered to be the main center of tomato (Solanum lycopersicum L.) domestication, since it has great genetic variation as reflected in the diverse traits found in native varieties (Peralta & Spoonner, 2007).
Knowledge of an ecosystem’s biodiversity is of great importance for the immediate conservation of genetic resources; this requires having an inventory to effectively quantify and conserve the genetic variation, in order to achieve sustainable use and exploitation through breeding (Benavides-González, Cisne-Contreras, Querol-Lipcovich, & Morán-Centeno, 2011). This leads to the need to characterize wild and native species through their morphology or molecular DNA markers.
Despite the importance of recognizing the characteristics and patterns of genetic variation in genetic resources, it is estimated that 80 % of the collections created worldwide have not been characterized and 95 % remain without agronomic evaluation; thus, the role of seedbanks is limited to just storing seed (Boada-Higuera, Mejía-Ramírez, Ceballos-Aguirre, & Orozco, 2010). On the other hand, with the growing number of accessions to conserve, it is essential to detect and remove duplicate materials and, additionally, discover the value of the collections conserved (González-Aguilera et al., 2011). This will facilitate their use through the rescue of traits of interest for breeding (Benavides-González et al., 2011).
Studies have been conducted in order to generate groups according to their genetic similarity or characteristics of anthropocentric interest, such as postharvest quality, fruit shapes and colors, culinary quality, nutraceutical quality, taste and agronomic benefits (Carrillo-Rodríguez & Chávez-Servia, 2010; Crisanto-Juárez, Vera-Guzmán, Chávez-Servia, & Carrillo-Rodríguez, 2010). The purpose of this research has been to generate information to identify sets of genotypes with high variation among them, so that their differentiation is evident; but at the same time, groups must be homogeneous in their interior.
Genetic variation in plant species can be explored in a fast, accurate and efficient manner by using molecular markers (González-Aguilera et al., 2011). Different molecular markers have been used in tomato, including ISSR (Robinson & Harris, 2000), which do not require prior knowledge of the genome and generate high polymorphism, facilitating studies of genetic diversity, phylogenetics, genomics and biological evolution (González-Aguilera et al., 2011).
Sequencing techniques have verified that there is very little polymorphism at the DNA level in commercial tomato (Park, West, & St-Clair, 2004). In the case of 96 tomato accessions characterized by González-Aguileraet al. (2011) with ISSR primers, they generated 144 bands, of which only 53 showed polymorphism, with an average of 14.4 bands per primer.
In this situation, the wild relatives of tomatoes are one of the main sources of germplasm for breeding this crop, since they have developed multiple traits that have allowed them to survive in adverse environmental conditions, including pests and diseases (Eingenbrode, Trumble, & Jones, 1993; Pérez-Grajales, Márquez-Sánchez, & Peña-Lomelí, 1997); consequently, efforts have been made to collect and characterize materials native to Mexico.
Based on the above, this research was conducted in order to evaluate the genetic variation in 55 native tomato (Solanum lycopersicum L.) collections from nine states of Mexico, through their morphological and molecular characterization, and create a strategy for their sustainable and efficient conservation.
Materials and methods
Collections evaluated
A total of 55 native tomato collections from nine states in the Mexican Republic were characterized morphologically and molecularly: Campeche (C14), Chiapas (C6, C7, C8, C24, C33, C36, C37, C50, C58, C59), Guanajuato (C43), Guerrero (C60), Hidalgo (C55), Jalisco (C39, C40, C47), Oaxaca (C4, C12, C17, C19, C20, C21, C22, C27, C29, C32, C45, C46, C61), Puebla (C2, C3, C16, C48, C49, C52, C56), San Luis Potosí (C11, C18), Tabasco (C1, C53), Veracruz (C5, C9, C25, C26, C31, C35, C42, C51). Additionally, eight collections of unknown origin (C15, C28, C54, C62, C63 and C64) were studied.
Morphological characterization
The morphological characterization was carried out in the 2013 spring-summer growing season in a full-vent, medium-technology greenhouse with 600-grade polyethylene covering with 70 % light transmission, and front, side and top ventilation protected by anti-aphid mesh, located at Chapingo Autonomous University (19° 29' NL and 98° 53' WL; 2240 masl), with a mean annual temperature of 15.9 °C. The collections were sown in 200-cavity polystyrene trays with peatmoss substrate.
Transplanting was performed 30 days after sowing; the plants were grown hydroponically in 15-kg polypropylene pots. The substrate used was volcanic pumice (tezontle) with particles of less than 3 mm in diameter. The nutrient solution used was that proposed by Cadahia-López (2000), and the amount applied varied according to the phenological stage and weather conditions. The conduction system was a stem and the planting density was 3.7 plants·m-2. The experimental unit consisted of a pot with two plants. A randomized complete block experimental design with five replications was used.
The morphological characterization was carried out by quantifying 62 qualitative and quantitative descriptors according to the IPGRI guide for tomato.
Molecular characterization
The molecular characterization was performed in 2014 at Chapingo Autonomous University’s Laboratory for Assisted Plant Breeding. DNA was extracted from the collections using the protocol described by Wagner et al. (1987). To determine the DNA concentration obtained in ng·μL-1, a Thermo Scientific NanoDrop Lite spectrometer was used. DNA quality assessment was performed on 0.8 % agarose gel by electrophoresis with 90 volts for 1.5 h. Subsequently the gel was stained in an ethidium bromide solution (0.6 μg·μL-1 in TAE 1 X) for 20 minutes. Then it was documented using a UVP-brand transilluminator and the UVP Doc-It LS Image Acquisition package.
Taq DNA polymerase chain reactions were performed with 16 primers (Table 1). For each genotype, 2.5 μL of DNA diluted to 10 ng·μL-1 were added to a 0.2-mL Eppendorf tube with 22.5 μL of reaction mixture; thus, the final volume was 25 μL. The reaction mixture consisted of: 5.2 μL of Molecular Biology grade H2O, 10 μL of dNTPs (500 μM), 2.5 μL of buffer (10 x), 1.5 μL of MgCl2 (50 mM), 3.0 μL of ISSR primer (10 ng·μL-1), 0.3 μL of Taq DNA polymerase enzyme (5 u·μL-1), and 2.5 μL of DNA (10 ng·μL-1). Amplification was carried out in a TECHNE-brand F TC41H2D model thermocycler, with the following program: a pre-denaturation cycle at a temperature of 93 °C for 1 min; forty denaturation cycles, each with: 20 s at 93 °C, 60 s at the annealing temperature of the primer used and 20 s at 72 °C; final extension at 72 °C for 6 min. Finally the sample was cooled to 10 °C.
Amplified products were separated by electrophoresis in agarose gel, stained with ethidium bromide and documented by a transilluminator and the UVP-brand Doc-It LS Image Acquisition package.
Encoding of the information obtained in the gels was carried out based on the similarities of the banding patterns by assigning the value 0 to the absence and 1 to the presence of each band. The number of polymorphic bands resulting from amplification was quantified for each primer, and each band was identified according to the migration distance in the gel.
Statistical analysis
For the analysis of morphological characters, the most useful variables for differentiating collections were selected by means of successive covariance analyses. Subsequently, cluster analysis was conducted from the squared Euclidean distance matrix and the generation algorithm of Ward’s minimum distance dendrogram; the height cutoff point was decided based on the cubic clustering criterion (Statistical Analysis System [SAS], 1983), the pseudo t2 statistic (Hotelling, 1951) and the pseudo F statistic (Johnson, 1998).
Table 1.
| Primer | Sequence | Annealing temperature °C |
| ISSR1 | (C A)8 A A G G | 62 |
| ISSR3 | (C A)8 A A G C T | 49 |
| ISSR4 | (A G)8 C T C | 55 |
| ISSR6 | (A C)8 C T G | 59 |
| ISSR7 | (A G)8 C T G | 53 |
| ISSR8 | (A C)8 C T T | 57 |
| ISSR9 | (A G)8 C | 47 |
| ISSR10 | (G A)8 T | 43 |
| UBC820 | (G T)8 C | 50 |
| UBC822 | (T C)8 A | 46 |
| UBC823 | (T C)8 C | 48 |
| UBC829 | (T G)8 C | 56 |
| UBC840 | (G A)8 C T T | 51 |
| UBC862 | (A G C)6 | 68 |
| UBC866 | (C T C)6 | 60 |
| LOL 7 | (G A)6 C C | 44 |
In order to corroborate the appropriateness of the grouping generated and identify the characters responsible for it, discriminant analysis in which the generated groups were considered as the categorical variable was performed. Additionally, resubstitution tests were conducted to verify the appropriateness of each collection to the assigned group (Johnson, 1998).
The molecular data were used to obtain the Jaccard (1912) index, with which a cluster analysis was performed with the methodology described above.
Results and discussion
Morphological characterization
Selection of informative variables
From 62 evaluated morphological characters, successive analyzes of covariance were able to reduce the dimensionality of the data matrix by identifying 15 with greater relevance for discriminating collections: seed weight, primary leaf length, primary leaf width, fruit shape, immature fruit color, fruit length, fruit width, fruit firmness, pericarp color, number of locules, sepal length, petal length, stamen length, plant size and number of nodes. These characters were used for the subsequent multivariate analyzes; the rest were not considered to be useful to differentiate collections, or because of their high association with the selected variables.
The above results indicate that all plant structures showed variability (leaves, fruits, flowers and plant size), which was useful for differentiating the collections. In a study by Pacheco-Triste, Chávez-Servia, and Carrillo-Rodríguez (2014), similar results where the differences among characterized populations were due to the divergence in fruit and plant characteristics were obtained; on the other hand, Carrillo-Rodríguez and Chávez-Servia (2010) found that the number of flowers per cluster, average fruit weight, fruits per cluster and total number of fruits to the fifth cluster had high variation among collections studied under greenhouse conditions.
Cluster analysis
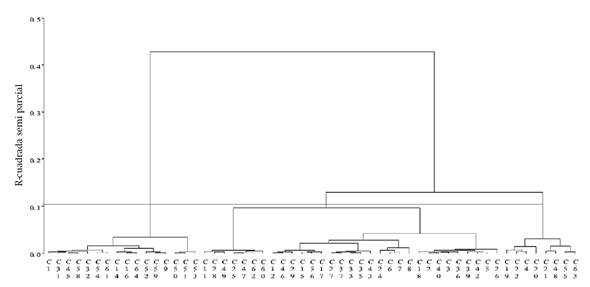
The dendrogram in Figure 1, built with Ward’s minimum variance algorithm, with a height cutoff point of semipartial 0.1 r2, defined from the cubic clustering criterion, pseudo F and pseudo t2, determined that the number of optimal groups was three, which included 16, 31 and 8 collections, respectively. While most studies use the UPGMA method, the present one considers that when working with collections from various regions of the country made with different criteria, Ward’s method is suitable for this type of analysis, based on the advantages mentioned by Núñez-Colín and Escobedo-López (2011): it reduces the presence of outliers and is most useful when the aim of the study is to determine the existing variability in genotypes of a specific species or genus.
The discriminant analysis, conducted in order to corroborate the groupings of the previous analysis and detect the relative importance of the characters in the formation of the three groups, considered these groupings as the categorical variable. Being three groups included in the analysis, the two discriminant variables (DV) generated explained the total variability observed. The first DV accounted for 80.7 % and the second the remaining 19.3 %.
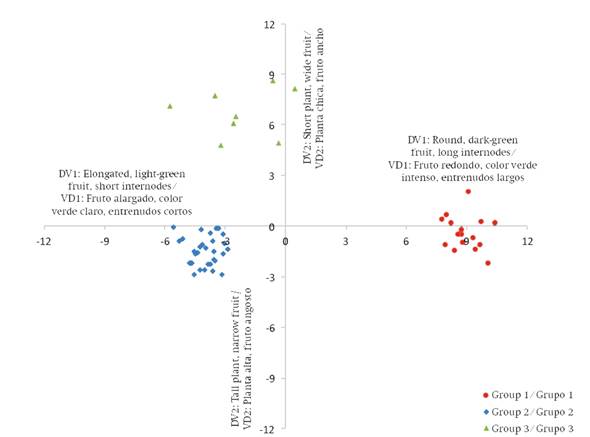
The total canonical structure indicated the linear associations between the discriminant functions generated and the 15 characters used in the analysis. DV1 was positively associated with fruit shape, fruit width and number of locules, and negatively with the number of nodes, which means that collections with high DV1 values had rounder fruits, a bright green color before maturity, greater fruit width and fewer nodes in a linear meter of plant and vice versa. With respect to DV2, it was positively associated with fruit length and width, and negatively with plant size, that is, collections with high values in DV2 have longer fruit and small size plant and vice versa. Based on the above, the morphological characteristics of the generated groups and the genotypes that comprise them are shown in Table 2.
Table 2.
| Group | Frequency | Collection | Characteristics |
|---|---|---|---|
| 1 | 16 | 1, 9, 14, 16, 31, 32, 45, 50, 51, 52, 53, 54, 58, 59, 61, 64 | Plants with few nodes, oblate fruits of medium size, soft green color before maturity and few locules per fruit |
| 2 | 31 | 11, 25, 28, 47, 49, 60, 62, 2, 3, 5, 6, 7, 8, 12, 15, 17, 18, 24, 26, 27, 29, 33, 35, 36, 37, 39, 40, 42, 43, 46,56 | Smaller-sized plants, with small, round fruits, dark green before maturity and a larger number of locules. |
| 3 | 8 | 4, 19, 20, 21, 22, 48, 55, 63 | Plants with few nodes, large cylindrical fruits, light green color before maturity and fewer locules. |
Similar results were found by Vásquez-Ortiz, Carrillo-Rodríguez, and Ramírez-Vallejo (2010) by differentiating collections based on qualitative and quantitative fruit, flower and leaf characteristics. A study by Agudelo-Agudelo, Ceballos-Aguirre, and Orozco (2011) showed that corolla color, hypocotyl pubescence and leaf type were not useful for differentiating collections, whereas fruit color, intensity and shape, along with the type of plant growth, were efficient in differentiating collections.
From the DV1 and DV2 values assigned to each collection, Figure 2 was generated. In it, the formation of groups obtained in the cluster analysis is verified.
Group 1 consisted of plants with kidney-type fruits, while group 2 was made up of collections with mostly cherry-type fruit, some kidney shaped and one cylindrical, which are similar in length and width, so that they resemble round fruits. Group 3 contains genotypes with large fruits by having the largest diameter, with cylindrical or kidney shape. This same pattern of grouping based on fruit characters was reported by Carrillo-Rodríguez and Chávez-Servia (2010); for example, in a group with predominantly kidney-type fruits, they also found some pear-shaped ones; in another where a high frequency of fruits with less size and weight were grouped together, there were variations in shape, from large kidney-type fruits to very small round ones with a diameter of 1 cm.
The average fruit equatorial diameter of the 55 collections studied was 3.81 cm, while the polar diameter was 3.29 cm and the average number of locules was 3.18. Comparison of these values with those obtained in previous studies on tomato diversity (Bonilla-Barrientos et al., 2014; Álvarez-Hernández, Cortés-Madrigal, & García-Ruíz, 2009) suggests that, like Lobato-Ortiz et al. (2012) and Chávez-Servia, Carrillo-Rodríguez, Vera- Guzmán, Rodríguez-Guzmán, and Lobato-Ortiz (2011), there is great variability in native populations, which is very useful for breeding programs, which is why their study and conservation takes on such relevance.
In the resubstitution test performed with the linear discriminant functions (Johnson, 1998), it was confirmed that the grouping was done correctly because no evaluated collection was placed in any group other than the original one.
Molecular characterization
The 16 ISSR primers used in the molecular characterization generated 118 bands, of which only 81 were polymorphic, with variation percentages between 10 and 90 % within the collections studied. Thus, on average seven bands per primer were generated, of which five were polymorphic, accounting for 68.7 % of total polymorphism. Based on the results of González-Aguilera et al. (2011), the polymorphism level obtained is suitable for characterizing and establishing the genetic differences of the native tomatoes evaluated. The ISSR1 primer had the highest number of polymorphic bands (12) and 70.6 % polymorphism. The primers with the greatest polymorphism in the present study were those with the amino acid sequence (GA)8 or (CA)8, results that partially match those of González-Aguilera et al. (2011) and Tikunov, Khrustaleva, and Karlov (2003).
The technique used is very useful for determining the genetic variability in autogamic plants like tomatoes, since both from the results here and those of González-Aguilera et al. (2011) and Tikunov et al. (2003), the polymorphism detected allowed classifying the evaluated collections adequately.
Cluster analysis
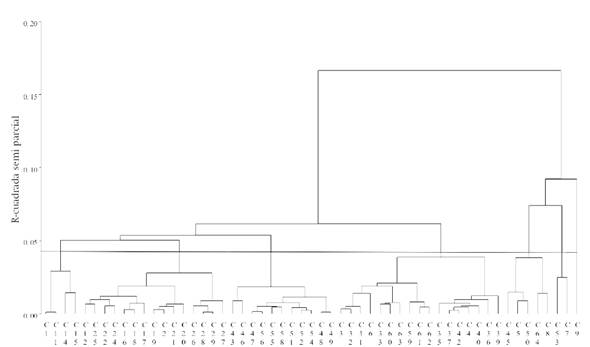
The Jaccard genetic distances, which averaged 0.48, along with the Ward method, generated the dendrogram in Figure 3. The height cutoff point (semipartial 0.04 r2), defined by the pseudo t2 and pseudo F, determined that the optimum number of groups was seven.
While most of the collections from the same region tend to cluster together, there are some collections from different areas within a specific group (Figure 3), so it is likely that they share adaptation areas with similar climatic and soil characteristics, which explains the greater similarity in their genetic constitution, a situation mentioned by Benor, Zhang, Wang, and Zhang (2008) in tomato lines where clusters were due to the geographical origin of the individuals.
Molecular and morphological characterization
When comparing the results of the groupings of the morphological and molecular characterizations as shown in Table 3, only 14 collections (26.5 %) followed the same grouping pattern and were placed within similar groups in the two analyzes. The other collections (73.5 %) were placed in different groups since in the morphological characters three groups were generated, while the molecular analysis generated seven.
Table 3.
| Molecular characterization groups | ||||
|---|---|---|---|---|
| 1 | 2 | 3 | ||
| Molecular characterization groups | 1 | 1R; 14R | 11CH; 15CH | - |
| 2 | 16R | 2CH; 12CH; 17CH; 18R; 24CH, 25CH; 26CH; 27CH; 28CH; 29CH | 19C; 20C; 21C | |
| 3 | 51C; 52R; 54R; 58R | 43CH; 46CH; 47CH; 49CH; 56CH | 48R; 55R | |
| 4 | 31R; 32R; 59R; 61R | 3CH; 6CH; 33CH; 35CH; 36CH; 37CH; 39CH; 40CH; 42CH; 60R; 62R | 4C; 22C; 63C | |
| 5 | 45R; 50R; 64R | 5CH; 8R | - | |
| 6 | 53R | 7CH | - | |
| 7 | 9R | - | - | |
The morphological characterization, by producing fewer groups, indicates that the characters used had less variation in the 55 collections; while the molecular characterization allowed us to more accurately estimate the genetic variation attributed to the ISSR markers, theoretically, they do not interact with the environment. On the other hand, by not expressing genetic variations restricted by the unique assessment environment (Boada-Higuera et al., 2010), morphological characterizations can lose accuracy in estimating the variation by confusing both the genetic effects and those of the genotype x environment interaction as suggested by Sultan (2000), who indicates that due to the phenotypic plasticity, a genotype can alter its development and physiology according to the environmental conditions where it grows.
Matches found in this study occurred only in group 2 of the morphological characterizations and group 2 of the molecular ones; the collections share the cherry-type fruit, and although they are not all from the same region of the country, they are native to places with similar environmental conditions, which determines the genetic similarity of these collections: C12 (Oaxaca, Cherry), C25 (Oaxaca, Cherry), C24 (Chiapas, Cherry), C18 (S.L.P., Kidney), C17 (Oaxaca, Cherry), C2 (Puebla, Cherry), C26 (Veracruz, Cherry), C28 (Chiapas, Cherry), C29 (Oaxaca, Cherry), C27 (Veracruz, Cherry). Benor et al. (2008) generated an initial grouping based on molecular matches; on a second level, subgroups of lines with similar origins, which associated with genetic similarities, were formed.
Although the collections were placed in morphological groups different from those generated by molecular groups, the results suggest that the ISSR markers are of great help in differentiating materials and determining the collections repeated in a seedbank, avoiding the storage of materials with the same characteristics that are not expressed in the same assessment environment (González-Aguilera et al., 2011; Zuriaga et al., 2009). For example, collections 19, 20 and 21 have the same type of fruit and are from the same place; thus, it is possible to make a decision on the collection to conserve based on the greater polymorphism obtained.
The morphological characterization lasted nine months under greenhouse conditions with 62 variables recorded, while the molecular characterization was performed in only three months based on plant material obtained from seedlings. Since they did not follow the same clustering pattern, performing both resulted in a 37.5 % increase in the efficiency of differentiating the genotypes, so they should be considered complementary and indispensable. This improves the ability to detect differences between genotypes and increases accuracy by generating conservation groups with greater genetic variability.
Conservation strategy
The results of the morphological characterization were due to a set of phenotypic variables, mainly associated with fruit shape; this allowed differentiating collections and placing them into three groups homogenous inside and heterogeneous among them (Group 1 kidneys, Group 2 cherry and Group 3 cylindrical or round). Choosing which genotypes to conserve within each set can be based on other qualities such as: origins, fruit color, fruit shape, anthropocentric use, etc.
By contrast, when there is only a molecular characterization, it is not possible to weight or differentiate the collections that make up a group, since the markers are random and dominant and the accuracy of the genetic variation estimate may be lost. Another consideration in this discussion is that collections are usually composed of several genotypes, so by using mixtures of DNA from the evaluated plants in order to save resources, the possibility of individually identifying them is lost; thus, the choosing of collections to conserve within each group would be random.
In the present study the morphological characterization detected less diversity than the molecular one and generated different clustering patterns, confirming that they are complementary studies as demonstrated by Demey, Zambrano, Fuenmayor, and Segovia (2003).
When both characterizations were combined, group 1 of the morphological characterization, with kidney-type fruits, was divided into seven subgroups by the molecular characterization, whereas group 2, with cherry-type fruits, was divided into 6 subgroups and group 3, with cylindrical fruits, into 3 subgroups. This generated 16 subgroups (Table 3).
Under these considerations, molecular characterization greatly improves the efficiency involved in conserving genetic resources since it allows estimating within each group obtained in the morphological characterization the genetic variation not detected by the former; thus, it decreases the classification error and allows adequately analyzing the genetic variability studied by both characterizations as mentioned by Demey et al. (2003).
According to Rao et al. (2007), seed samples to be conserved should be genetically distinct from any other accession already registered in a seedbank. In the present situation this can be ensured by conserving one collection of each of the 16 subgroups identified in Table 3, or by a mixture of seeds of the genotypes comprising each subgroup.
Based on the above, in order to efficiently conserve the genetic variation studied, the selection of the genotypes of each subgroup (Table 3), for their safekeeping in the seedbank, will be based on maintaining different origins and differential fruit characters such as color and shape. If the number of collections is not reduced with the above criteria, the last selection criterion will be anthropocentric use.
Thus, the proposal for conserving the genetic variation in the 55 collections evaluated, and based on the criteria mentioned above, consists of safeguarding the 37 collections shown in Table 3, representing 67 % of the total number of collections evaluated, thereby eliminating genotype duplication.
In a more dramatic case, and based on the fact that the DNA characterized is a mixture from several individuals that make up a collection, the recommendation would be to maintain a balanced seed sample of the collections comprising a subgroup. Thus, in our case 16 samples would be conserved.
Conclusions
This research provides a basis for the efficient conservation of collections by avoiding the multiplicity of materials with similar characters.
The morphological and molecular characterizations are complementary in that together they more accurately assess the genetic variability of the populations studied, enabling a sustainable and more efficient conservation of the phytogenetic resources of tomato.
The evaluated collections from eight states of Mexico showed wide genetic variation identified through 16 conservation groups, which can be conserved by safeguarding 67 % of the collections studied.
